20.1 Organizing Life

This bee and Echinacea flower (Figure 20.1) could not look more different, yet they are related, as are all living organisms on Earth. By following pathways of similarities and changes—both visible and genetic—scientists seek to map the evolutionary past of how life developed from single-celled organisms to the tremendous collection of creatures that have germinated, crawled, floated, swum, flown, and walked on this planet.
Learning Outcomes
- Discuss the need for a comprehensive classification system
- List the different levels of the taxonomic classification system
- Describe how systematics and taxonomy relate to phylogeny
- Discuss a phylogenetic tree’s components and purpose
In scientific terms, phylogeny is the evolutionary history and relationship of an organism or group of organisms. A phylogeny describes the organism’s relationships, such as from which organisms it may have evolved, or to which species it is most closely related. Phylogenetic relationships provide information on shared ancestry but not necessarily on how organisms are similar or different.
Phylogenetic Trees
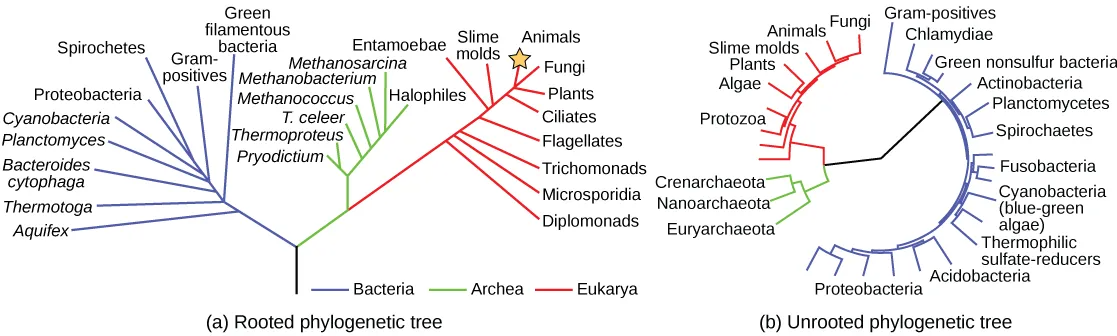
Scientists use a tool called a phylogenetic tree to show the evolutionary pathways and connections among organisms. A phylogenetic tree is a diagram used to reflect evolutionary relationships among organisms or groups of organisms. Scientists consider phylogenetic trees to be a hypothesis of the evolutionary past since one cannot go back to confirm the proposed relationships. In other words, we can construct a “tree of life” to illustrate when different organisms evolved and to show the relationships among different organisms (Figure 20.2).
Unlike a taxonomic classification diagram, we can read a phylogenetic tree like a map of evolutionary history. Many phylogenetic trees have a single lineage at the base representing a common ancestor. Scientists call such trees rooted, which means there is a single ancestral lineage (typically drawn from the bottom or left) to which all organisms represented in the diagram relate. Notice in the rooted phylogenetic tree that the three domains— Bacteria, Archaea, and Eukarya—diverge from a single point and branch off. The small branch that plants and animals (including humans) occupy in this diagram shows how recent and miniscule these groups are compared with other organisms. Unrooted trees do not show a common ancestor but do show relationships among species.
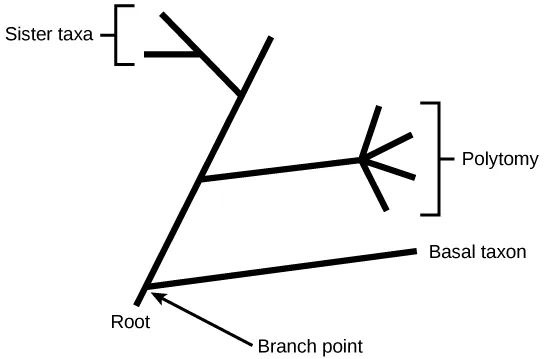
In a rooted tree, the branching indicates evolutionary relationships (Figure 20.3). The point where a split occurs, a branch point, represents where a single lineage evolved into a distinct new one. We call a lineage that evolved early from the root that remains unbranched a basal taxon. We call two lineages stemming from the same branch point sister taxa. A branch with more than two lineages is a polytomy and serves to illustrate where scientists have not definitively determined all of the relationships. Note that although sister taxa and polytomy do share an ancestor, it does not mean that the groups of organisms split or evolved from each other. Organisms in two taxa may have split at a specific branch point, but neither taxon gave rise to the other.
The diagrams above can serve as a pathway to understanding evolutionary history. We can trace the pathway from the origin of life to any individual species by navigating through the evolutionary branches between the two points. Also, by starting with a single species and tracing back towards the “trunk” of the tree, one can discover species’ ancestors, as well as where lineages share a common ancestry. In addition, we can use the tree to study entire groups of organisms.
Another point to mention on phylogenetic tree structure is that rotation at branch points does not change the information. For example, if a branch point rotated and the taxon order changed, this would not alter the information because each taxon’s evolution from the branch point was independent of the other.
Many disciplines within the study of biology contribute to understanding how past and present life evolved over time; these disciplines together contribute to building, updating, and maintaining the “tree of life.” Systematics is the field that scientists use to organize and classify organisms based on evolutionary relationships. Researchers may use data from fossils, from studying the body part structures, or molecules that an organism uses, and DNA analysis. By combining data from many sources, scientists can construct an organism’s phylogeny. Since phylogenetic trees are hypotheses, they will continue to change as researchers discover new types of life and learn new information.
Limitations of Phylogenetic Trees
It may be easy to assume that more closely related organisms look more alike, and while this is often the case, it is not always true. If two closely related lineages evolved under significantly varied surroundings, it is possible for the two groups to appear more different than other groups that are not as closely related. For example, the phylogenetic tree in Figure 20.4 shows that lizards and rabbits both have amniotic eggs; whereas, frogs do not. Yet lizards and frogs appear more similar than lizards and rabbits.
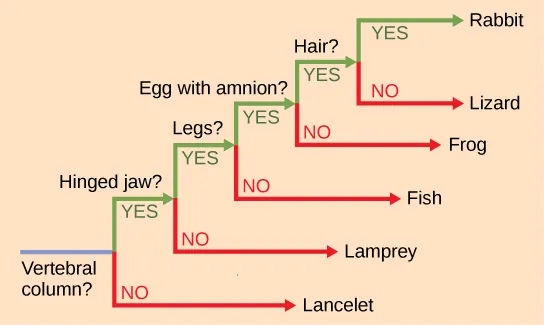
Another aspect of phylogenetic trees is that, unless otherwise indicated, the branches do not account for length of time, only the evolutionary order. In other words, a branch’s length does not typically mean more time passed, nor does a short branch mean less time passed— unless specified on the diagram. For example, in Figure 20.4, the tree does not indicate how much time passed between the evolution of amniotic eggs and hair. What the tree does show is the order in which things took place. Again using Figure 20.4 the tree shows that the oldest trait is the vertebral column, followed by hinged jaws, and so forth. Remember that any phylogenetic tree is a part of the greater whole, and like a real tree, it does not grow in only one direction after a new branch develops. Thus, for the organisms in Figure 20.4, just because a vertebral column evolved does not mean that invertebrate evolution ceased. It only means that a new branch formed. Also, groups that are not closely related, but evolve under similar conditions, may appear more phenotypically similar to each other than to a close relative.
Link to Learning
Building phylogenetic trees based on homologies
Head to this website to use interactive exercises that allow you to explore the evolutionary relationships among species by building your own phylogenetic trees based on species morphological characteristics and their DNA sequences. Working through the exercises, you will understand better how phylogenetic trees are built, and how to “read” them.
Click on the “PLAY GAME” button to start working through the puzzles. It will take you some time (~1.5 hours) to go through all the tree building missions, and you might find this a challenging exercise at first! However, as your understanding of phylogenies grows as we progress through this module, you might find the puzzles become easier. So, feel free to come back to this exercise throughout this module to continue. If you want to save your progress in the meantime, you will have to create an account on the website. Otherwise, you can just play as a guest.
Classification Levels
Taxonomy (which literally means “arrangement law”) is the science of classifying organisms to construct internationally shared classification systems with each organism placed into increasingly more inclusive groupings. Think about a grocery store’s organization. One large space is divided into departments, such as produce, dairy, and meats. Then each department further divides into aisles, then each aisle into categories and brands, and then finally a single product. We call this organization from larger to smaller, more specific categories a hierarchical system.
The taxonomic classification system (also called the Linnaean system after its inventor, Carl Linnaeus, a Swedish botanist, zoologist, and physician) uses a hierarchical model. Moving from the point of origin, the groups become more specific, until one branch ends as a single species. For example, after the common beginning of all life, scientists divide organisms into three large categories called domains: Bacteria, Archaea, and Eukarya. Within each domain is a second category called a kingdom. After kingdoms, the subsequent categories of increasing specificity are: phylum, class, order, family, genus, and species (Figure 20.5).
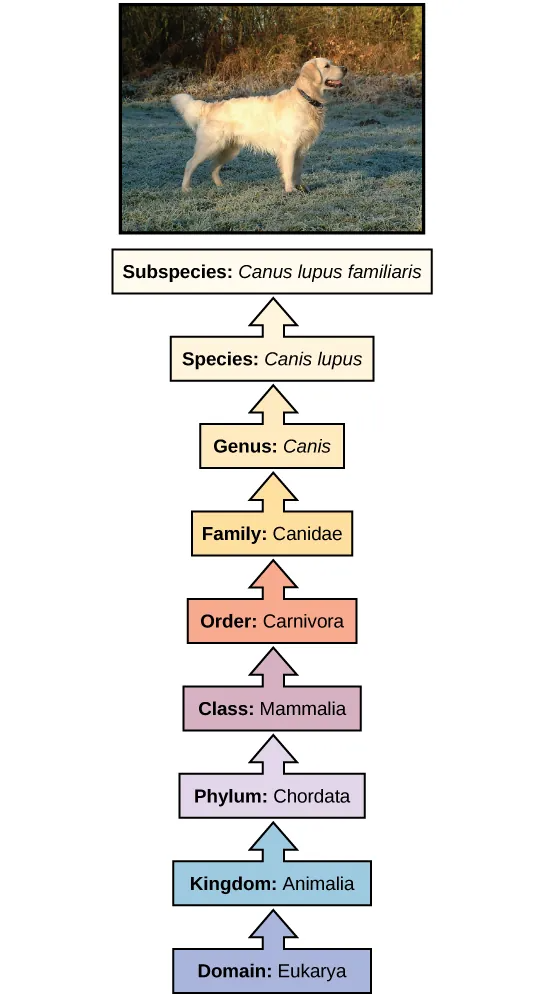
The kingdom Animalia stems from the Eukarya domain. Figure 20.5 above shows the classification for the common dog. Therefore, the full name of an organism technically has eight terms. For the dog it is: Eukarya, Animalia, Chordata, Mammalia, Carnivora, Canidae, Canis, and lupus. Notice that each name is capitalized except for species, and the genus and species names are italicized. Scientists generally refer to an organism only by its genus and species, which is its two-word scientific name, or binomial nomenclature. Therefore, the scientific name of the dog is Canis lupus. The name at each level is also a taxon. In other words, dogs are in order Carnivora. Carnivora is the name of the taxon at the order level; Canidae is the taxon at the family level, and so forth. Organisms also have a common name that people typically use, in this case, dog. Note that the dog is additionally a subspecies: the “familiaris” in Canis lupus familiaris. Subspecies are members of the same species that are capable of mating and reproducing viable offspring, but they are separate subspecies due to geographic or behavioral isolation or other factors.
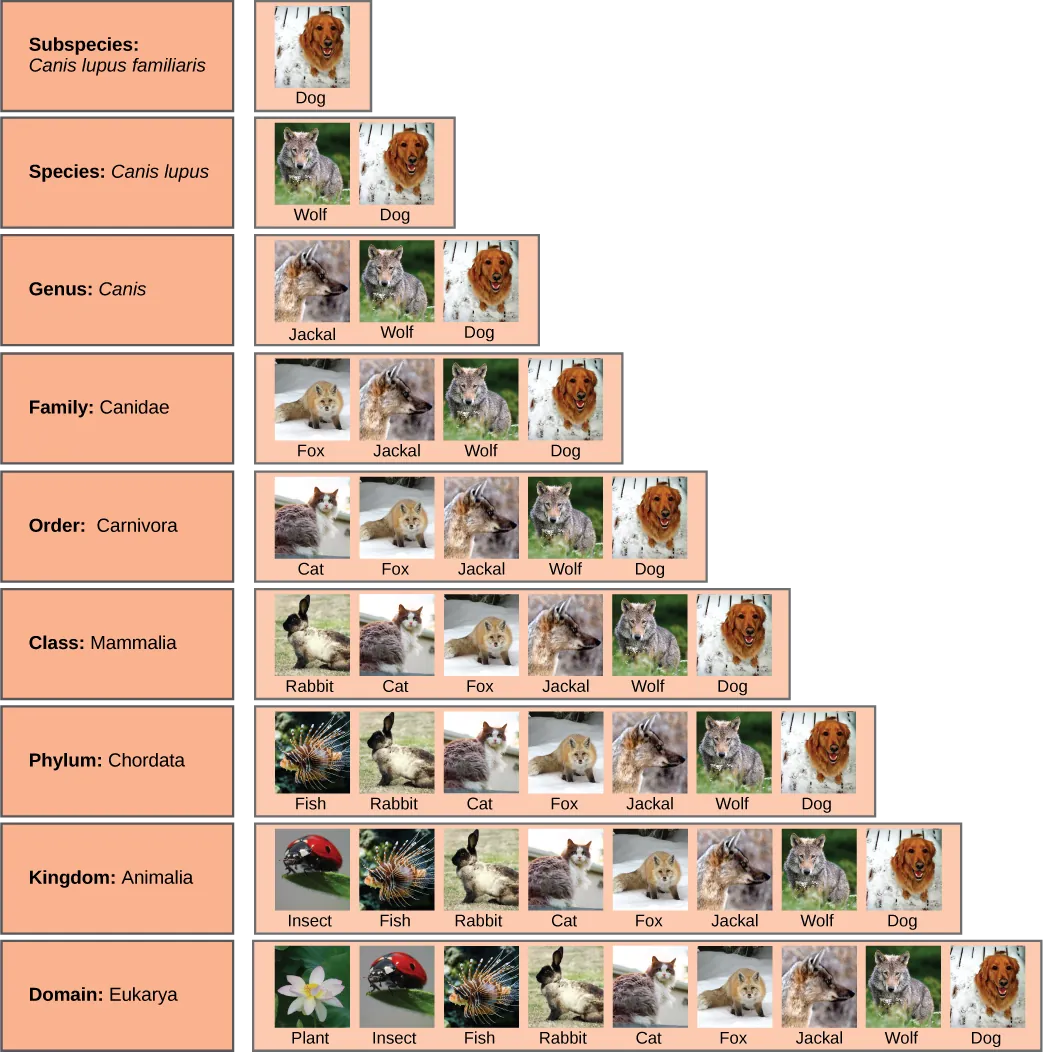
Figure 20.6 shows how the levels move toward specificity with other organisms. Notice how the dog shares a domain with the widest diversity of organisms, including plants and butterflies. At each sublevel, the organisms become more similar because they are more closely related. Historically, scientists classified organisms using characteristics, but as DNA technology developed, they have determined more precise phylogenies.
Link to Learning
Taxonomy
This video by Crash Course recaps the concept of taxonomy.
Table of Contents
- Taxonomy 0:00
- Phylogenetic Tree 1:24
- Biolography 2:26
- Analogous/Homoplasic Traits 3:48
- Homologous Traits 4:03
- Taxa & Binomial Nomenclature 4:56
- Domains 5:48
- Bateria 6:04
- Archaea 6:44
- Eukarya / 4 Kingdoms 6:54
- Plantae 7:56
- Protista 8:23
- Fungi 8:56
- Animalia 9:31
Recent genetic analysis and other advancements have found that some earlier phylogenetic classifications do not align with the evolutionary past; therefore, researchers must make changes and updates as new discoveries occur. Recall that phylogenetic trees are hypotheses and are modified as data becomes available. In addition, classification historically has focused on grouping organisms mainly by shared characteristics and does not necessarily illustrate how the various groups relate to each other from an evolutionary perspective. For example, despite the fact that a hippopotamus resembles a pig more than a whale, the hippopotamus may be the whale’s closest living relative.
evolutionary history and relationship of an organism or group of organisms
diagram showing the evolutionary relationships among various biological species based on similarities and differences in genetic or physical traits or both; in essence, a hypothesis concerning evolutionary connections
single ancestral lineage on a phylogenetic tree to which all organisms represented in the diagram relate
node on a phylogenetic tree where a single lineage splits into distinct new ones
branch on a phylogenetic tree that has not diverged significantly from the root ancestor
two lineages that diverged from the same branch point
branch on a phylogenetic tree with more than two groups or taxa
field of organizing and classifying organisms based on evolutionary relationships
science of classifying organisms
system of two-part scientific names for an organism, which includes genus and species names
(plural: taxa) single level in the taxonomic classification system

